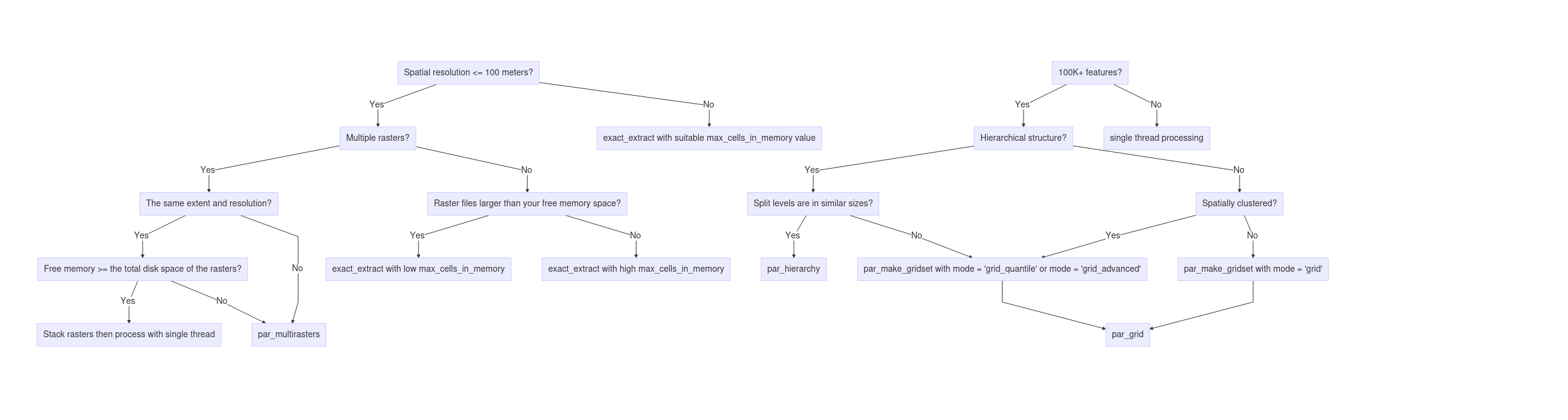
Examples
- Examples will navigate
par_grid, par_hierarchy, and par_multirasters functions in chopin to parallelize geospatial operations.
# check and install packages to run examples
pkgs <- c("chopin", "dplyr", "sf", "terra",
"future", "future.apply", "doFuture", "testthat")
# install packages if anything is unavailable
rlang::check_installed(pkgs)
# disable spherical geometries
# it does the same as sf::sf_use_s2(FALSE)
options(sf_use_s2 = FALSE)
# parallelization-safe random number generator
set.seed(2024, kind = "L'Ecuyer-CMRG")
par_grid: parallelize over artificial grid polygons
- Please refer to a small example below for extracting mean altitude values at circular point buffers and census tracts in North Carolina.
- Before running code chunks below, set the cloned
chopin repository as your working directory with setwd()
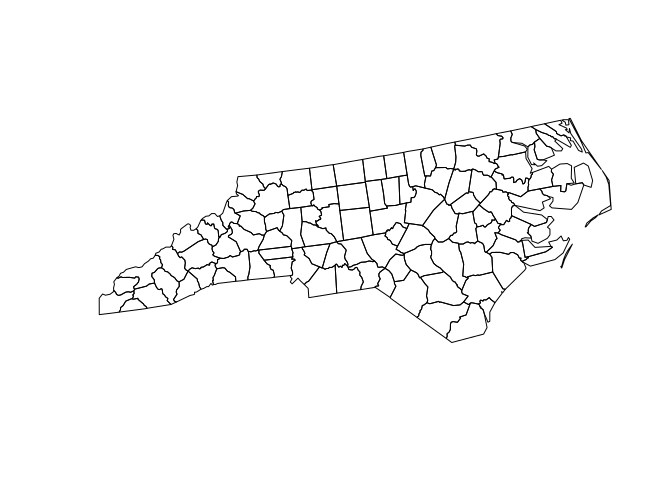
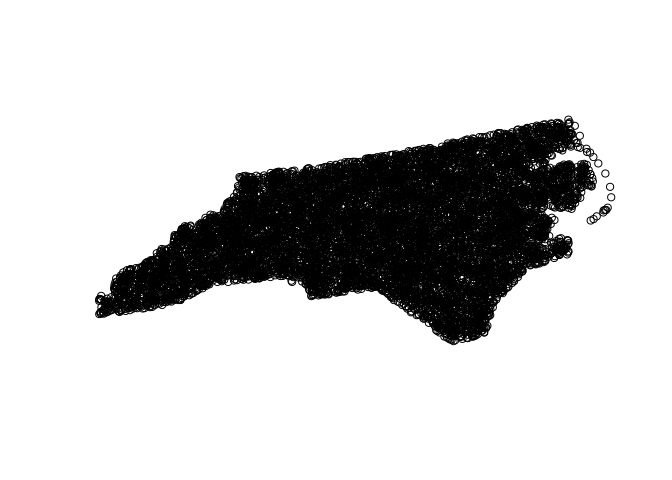
Generate random points in NC
- Ten thousands random point locations were generated inside the counties of North Carolina.

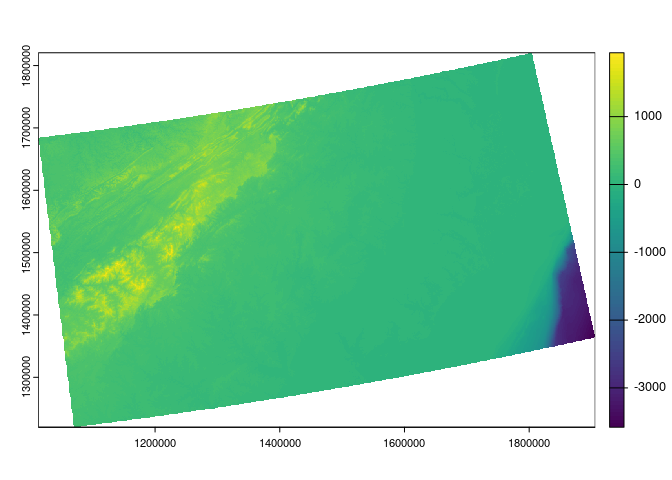
- We use an elevation dataset with and a moderate spatial resolution (approximately 400 meters or 0.25 miles).
# data preparation
wdir <- system.file("extdata", package = "chopin")
path_srtm <- file.path(wdir, "nc_srtm15_otm.rds")
# terra SpatRaster objects are wrapped when exported to rds file
srtm <- terra::unwrap(readRDS(path_srtm))
terra::crs(srtm) <- "EPSG:5070"
srtm
#> class : SpatRaster
#> dimensions : 1534, 2281, 1 (nrow, ncol, nlyr)
#> resolution : 391.5026, 391.5026 (x, y)
#> extent : 1012872, 1905890, 1219961, 1820526 (xmin, xmax, ymin, ymax)
#> coord. ref. : NAD83 / Conus Albers (EPSG:5070)
#> source(s) : memory
#> name : file928c3830468b
#> min value : -3589.291
#> max value : 1946.400
ncpoints_tr <- terra::vect(ncpoints)
system.time(
ncpoints_srtm <-
chopin::extract_at(
vector = ncpoints_tr,
raster = srtm,
id = "pid",
mode = "buffer",
radius = 1e4L # 10,000 meters (10 km)
)
)
#> user system elapsed
#> 9.925 0.197 10.149
Generate regular grid computational regions
-
chopin::par_make_gridset takes a spatial dataset to generate regular grid polygons with nx and ny arguments with padding. Users will have both overlapping (by the degree of radius) and non-overlapping grids, both of which will be utilized to split locations and target datasets into sub-datasets for efficient processing.
compregions <-
chopin::par_make_gridset(
ncpoints_tr,
mode = "grid",
nx = 8L,
ny = 5L,
padding = 1e4L
)
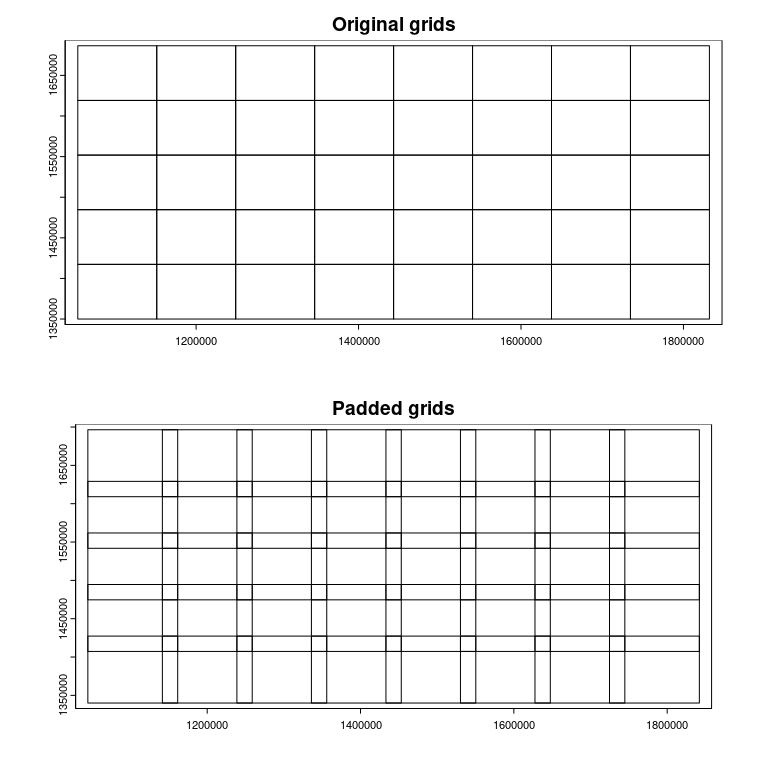
-
compregions is a list object with two elements named original (non-overlapping grid polygons) and padded (overlapping by padding). The figures below illustrate the grid polygons with and without overlaps.
names(compregions)
#> [1] "original" "padded"
oldpar <- par()
par(mfrow = c(2, 1))
terra::plot(compregions$original, main = "Original grids")
terra::plot(compregions$padded, main = "Padded grids")
Parallel processing
- Using the grid polygons, we distribute the task of averaging elevations at 10,000 circular buffer polygons, which are generated from the random locations, with 10 kilometers radius by
chopin::par_grid.
- Users always need to register multiple CPU threads (logical cores) for parallelization.
-
chopin::par_* functions are flexible in terms of supporting generic spatial operations in sf and terra, especially where two datasets are involved.
- Users can inject generic functions’ arguments (parameters) by writing them in the ellipsis (
...) arguments, as demonstrated below:
future::plan(future::multicore, workers = 4L)
system.time(
ncpoints_srtm_mthr <-
chopin::par_grid(
grids = compregions,
grid_target_id = NULL,
fun_dist = chopin::extract_at,
vector = ncpoints_tr,
raster = srtm,
id = "pid",
mode = "buffer",
radius = 1e4L
)
)
#> Your input function was successfully run at CGRIDID: 4
#> Your input function was successfully run at CGRIDID: 5
#> Warning: [buffer] empty SpatVector
#> Warning: [buffer] empty SpatVector
#> Warning: [buffer] empty SpatVector
#> Your input function was successfully run at CGRIDID: 6
#> Your input function was successfully run at CGRIDID: 7
#> Your input function was successfully run at CGRIDID: 8
#> Your input function was successfully run at CGRIDID: 10
#> Warning: [buffer] empty SpatVector
#> Your input function was successfully run at CGRIDID: 11
#> Your input function was successfully run at CGRIDID: 12
#> Your input function was successfully run at CGRIDID: 13
#> Your input function was successfully run at CGRIDID: 14
#> Your input function was successfully run at CGRIDID: 15
#> Your input function was successfully run at CGRIDID: 16
#> Your input function was successfully run at CGRIDID: 17
#> Your input function was successfully run at CGRIDID: 18
#> Your input function was successfully run at CGRIDID: 19
#> Your input function was successfully run at CGRIDID: 20
#> Your input function was successfully run at CGRIDID: 21
#> Your input function was successfully run at CGRIDID: 22
#> Your input function was successfully run at CGRIDID: 23
#> Your input function was successfully run at CGRIDID: 24
#> Your input function was successfully run at CGRIDID: 25
#> Your input function was successfully run at CGRIDID: 26
#> Your input function was successfully run at CGRIDID: 27
#> Your input function was successfully run at CGRIDID: 28
#> Your input function was successfully run at CGRIDID: 29
#> Your input function was successfully run at CGRIDID: 30
#> Your input function was successfully run at CGRIDID: 31
#> Your input function was successfully run at CGRIDID: 32
#> Your input function was successfully run at CGRIDID: 33
#> Your input function was successfully run at CGRIDID: 34
#> Warning: [buffer] empty SpatVector
#> Your input function was successfully run at CGRIDID: 37
#> Your input function was successfully run at CGRIDID: 38
#> Your input function was successfully run at CGRIDID: 39
#> Warning: [buffer] empty SpatVector
#> Warning: [buffer] empty SpatVector
#> user system elapsed
#> 8.418 1.567 3.447
colnames(ncpoints_srtm_mthr)[2] <- "mean_par"
ncpoints_compar <- merge(ncpoints_srtm, ncpoints_srtm_mthr)
# Are the calculations equal?
all.equal(ncpoints_compar$mean, ncpoints_compar$mean_par)
#> [1] TRUE
ncpoints_s <-
merge(ncpoints, ncpoints_srtm)
ncpoints_m <-
merge(ncpoints, ncpoints_srtm_mthr)
plot(ncpoints_s[, "mean"], main = "Single-thread", pch = 19, cex = 0.33)
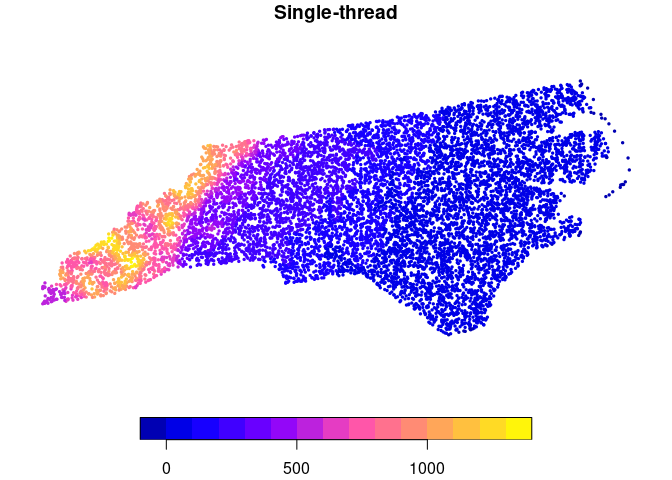
plot(ncpoints_m[, "mean_par"], main = "Multi-thread", pch = 19, cex = 0.33)
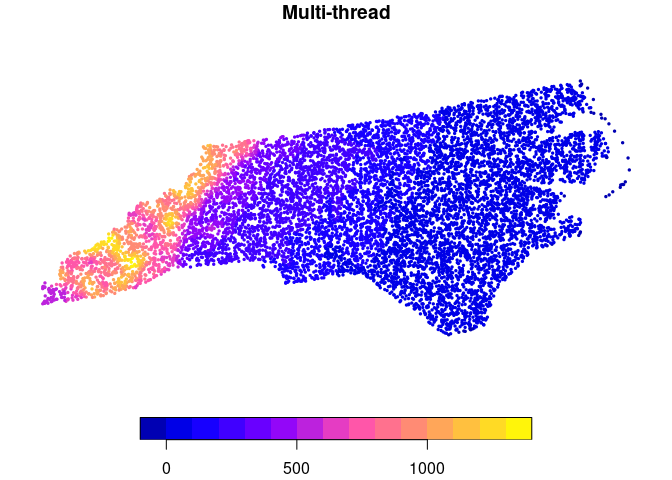
chopin::par_hierarchy: parallelize geospatial computations using intrinsic data hierarchy
- In real world datasets, we usually have nested/exhaustive hierarchies. For example, land is organized by administrative/jurisdictional borders where multiple levels exist. In the U.S. context, a state consists of several counties, counties are split into census tracts, and they have a group of block groups.
-
chopin::par_hierarchy leverages such hierarchies to parallelize geospatial operations, which means that a group of lower-level geographic units in a higher-level geography is assigned to a process.
- A demonstration below shows that census tracts are grouped by their counties then each county will be processed in a CPU thread.
Read data
path_nchrchy <- file.path(wdir, "nc_hierarchy.gpkg")
nc_data <- path_nchrchy
nc_county <- sf::st_read(nc_data, layer = "county")
#> Reading layer `county' from data source
#> `/tmp/RtmpdYJu7A/temp_libpath16ff5d6da0d43d/chopin/extdata/nc_hierarchy.gpkg'
#> using driver `GPKG'
#> Simple feature collection with 100 features and 1 field
#> Geometry type: POLYGON
#> Dimension: XY
#> Bounding box: xmin: 1054155 ymin: 1341756 xmax: 1838923 ymax: 1690176
#> Projected CRS: NAD83 / Conus Albers
nc_tracts <- sf::st_read(nc_data, layer = "tracts")
#> Reading layer `tracts' from data source
#> `/tmp/RtmpdYJu7A/temp_libpath16ff5d6da0d43d/chopin/extdata/nc_hierarchy.gpkg'
#> using driver `GPKG'
#> Simple feature collection with 2672 features and 1 field
#> Geometry type: MULTIPOLYGON
#> Dimension: XY
#> Bounding box: xmin: 1054155 ymin: 1341756 xmax: 1838923 ymax: 1690176
#> Projected CRS: NAD83 / Conus Albers
# reproject to Conus Albers Equal Area
nc_county <- sf::st_transform(nc_county, "EPSG:5070")
nc_tracts <- sf::st_transform(nc_tracts, "EPSG:5070")
nc_tracts$COUNTY <- substr(nc_tracts$GEOID, 1, 5)
Extract average SRTM elevations by single and multiple threads
# single-thread
system.time(
nc_elev_tr_single <-
chopin::extract_at(
vector = nc_tracts,
raster = srtm,
id = "GEOID",
mode = "polygon"
)
)
#> user system elapsed
#> 1.798 0.082 1.886
# hierarchical parallelization
system.time(
nc_elev_tr_distr <-
chopin::par_hierarchy(
regions = nc_county, # higher level geometry
regions_id = "GEOID", # higher level unique id
fun_dist = chopin::extract_at,
vector = nc_tracts, # lower level geometry
raster = srtm,
id = "GEOID", # lower level unique id
func = "mean"
)
)
#> user system elapsed
#> 10.452 2.020 4.324
par_multirasters: parallelize over multiple rasters
- There is a common case of having a large group of raster files at which the same operation should be performed.
-
chopin::par_multirasters is for such cases. An example below demonstrates where we have five elevation raster files to calculate the average elevation at counties in North Carolina.
nccnty <- terra::vect(nc_data, layer = "county")
ncelev <- terra::unwrap(readRDS(path_srtm))
terra::crs(ncelev) <- "EPSG:5070"
names(ncelev) <- c("srtm15")
tdir <- tempdir()
terra::writeRaster(ncelev, file.path(tdir, "test1.tif"), overwrite = TRUE)
terra::writeRaster(ncelev, file.path(tdir, "test2.tif"), overwrite = TRUE)
terra::writeRaster(ncelev, file.path(tdir, "test3.tif"), overwrite = TRUE)
terra::writeRaster(ncelev, file.path(tdir, "test4.tif"), overwrite = TRUE)
terra::writeRaster(ncelev, file.path(tdir, "test5.tif"), overwrite = TRUE)
# check if the raster files were exported as expected
testfiles <- list.files(tdir, pattern = "*.tif$", full.names = TRUE)
testfiles
#> [1] "/tmp/RtmpPWsyT6/test1.tif" "/tmp/RtmpPWsyT6/test2.tif"
#> [3] "/tmp/RtmpPWsyT6/test3.tif" "/tmp/RtmpPWsyT6/test4.tif"
#> [5] "/tmp/RtmpPWsyT6/test5.tif"
system.time(
res <-
chopin::par_multirasters(
filenames = testfiles,
fun_dist = chopin::extract_at_poly,
polys = nccnty,
surf = ncelev,
id = "GEOID",
func = "mean"
)
)
#> user system elapsed
#> 1.554 0.633 1.027
| 37037 |
136.80203 |
/tmp/RtmpPWsyT6/test1.tif |
| 37001 |
189.76170 |
/tmp/RtmpPWsyT6/test1.tif |
| 37057 |
231.16968 |
/tmp/RtmpPWsyT6/test1.tif |
| 37069 |
98.03845 |
/tmp/RtmpPWsyT6/test1.tif |
| 37155 |
41.23463 |
/tmp/RtmpPWsyT6/test1.tif |
| 37109 |
270.96933 |
/tmp/RtmpPWsyT6/test1.tif |
# remove temporary raster files
file.remove(testfiles)
#> [1] TRUE TRUE TRUE TRUE TRUE
Parallelization of a generic geospatial operation
- Other than
chopin internal macros, chopin::par_* functions support generic geospatial operations.
- An example below uses
terra::nearest, which gets the nearest feature’s attributes, inside chopin::par_grid.
path_ncrd1 <- file.path(wdir, "ncroads_first.gpkg")
# Generate 5000 random points
pnts <- sf::st_sample(ncsf, 5000)
pnts <- sf::st_as_sf(pnts)
# assign identifiers
pnts$pid <- sprintf("RPID-%04d", seq(1, 5000))
pnts <- terra::vect(pnts)
rd1 <- terra::vect(path_ncrd1)
# reproject
pnts <- terra::project(pnts, "EPSG:5070")
rd1 <- terra::project(rd1, "EPSG:5070")
# generate grids
nccompreg <-
chopin::par_make_gridset(
input = pnts,
mode = "grid",
nx = 4L,
ny = 2L,
padding = 5e4L
)
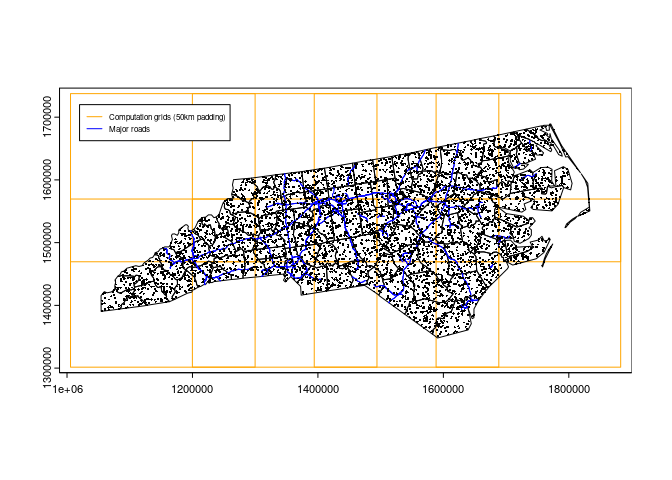
- The figure below shows the padded grids (50 kilometers), primary roads, and points. Primary roads will be selected by a padded grid per iteration and used to calculate the distance from each point to the nearest primary road. Padded grids and their overlapping areas will look different according to
padding argument in chopin::par_make_gridset.
# plot
terra::plot(nccompreg$padded, border = "orange")
terra::plot(terra::vect(ncsf), add = TRUE)
terra::plot(rd1, col = "blue", add = TRUE)
terra::plot(pnts, add = TRUE, cex = 0.3)
legend(1.02e6, 1.72e6,
legend = c("Computation grids (50km padding)", "Major roads"),
lty = 1, lwd = 1, col = c("orange", "blue"),
cex = 0.5)
# terra::nearest run
system.time(
restr <- terra::nearest(x = pnts, y = rd1)
)
#> user system elapsed
#> 0.462 0.003 0.466
# we use four threads that were configured above
system.time(
res <-
chopin::par_grid(
grids = nccompreg,
fun_dist = terra::nearest,
x = pnts,
y = rd1
)
)
#> Your input function was successfully run at CGRIDID: 1
#> Your input function was successfully run at CGRIDID: 2
#> Your input function was successfully run at CGRIDID: 3
#> Your input function was successfully run at CGRIDID: 4
#> Your input function was successfully run at CGRIDID: 5
#> Your input function was successfully run at CGRIDID: 6
#> Your input function was successfully run at CGRIDID: 7
#> Your input function was successfully run at CGRIDID: 8
#> user system elapsed
#> 0.776 0.562 0.521
- We will compare the results from the single-thread and multi-thread calculation.
resj <- merge(restr, res, by = c("from_x", "from_y"))
all.equal(resj$distance.x, resj$distance.y)
#> [1] TRUE
- Users should be mindful of potential caveats in the parallelization of nearest feature search, which may result in no or excess distance depending on the distribution of the target dataset to which the nearest feature is searched.
- For example, when one wants to calculate the nearest interstate from rural homes with fine grids, some grids may have no interstates then homes in such grids will not get any distance to the nearest interstate.
- Such problems can be avoided by choosing
nx, ny, and padding values in par_make_gridset meticulously.